In today’s post I wanted to describe classification trees. I will concentrate on Classification and Regression Tree (CART) algorithm. As I discuss the main features of this algorithm I will contrast it with other widely used methods to construct classification trees.
What A Classification Tree Looks Like:
Before diving into the algorithm let’s have a look at what a classification tree looks like. Let’s say we have a sample of the passenger list of the Titanic voyage that sunk in 1912, and based on some set of descriptive features of each passenger we want to be able to classify them as either having survived or not. This is a two class classification problem. For our features we have:
1) pcalss: information on which class each passenger was in during the journey. There were 1st, 2nd, and 3rd class passengers (I omit the crew in this model).
2) sex: the sex of each passenger
3) sibsp: number of siblings or number of spouses aboard
4) parch: number of parents or children aboard
An example of a classification tree that can be generated with this data is presented below:
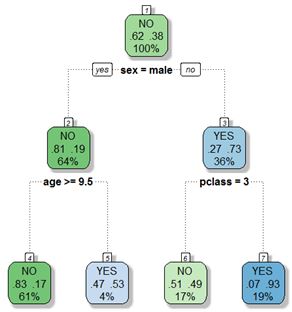
Trees have a root node at the top of the tree, leaf (or terminal nodes) at the bottom, with all other nodes being called intermediate nodes. To understand the model we follow along a path to a terminal node to see what the classification of a particular case might be. For example, the root node has 100% of the data with majority of observations (62% of passengers) not surviving the sinking of the ship. If a passenger was female then we move to the right on the classification tree to node labeled 3. In this node we can see that 73% of people survived. Furthermore, if this passenger was in 1st or 2nd class we would again move to the right and down the tree. In node 7 we can see that 93% of those passengers survived. However, if the female passenger was in 3rd class then she likely perished since 51% of the passengers in node 6 did not survive.
In the case of men, we move to node 2 from the root node. In this node, 81% of the individuals died. Furthermore, if the male was above the age of 9.5 then 83% of those individuals perished (node 4). Finally, if the male was below the age of 9.5 then it is likely that he survived (53% of the individuals in node 5 survived).
You can see that a tree can be described as a group of nested if statement. For example:
IF (male = TRUE & age >9.5) THEN Survive = NO – path from node 1 to node 4
IF (male = FALSE & class = 1 or 2) THEN Survive = YES – path from node 1 to node 7
I hope the reader can agree that there is something that is intuitive about how a tree classifies data. Trees are very appealing precisely because they can be easily interpreted.
Tree Growing Process:
Next I want to discuss how we can construct a classification trees. I will concentrate on the CART algorithm here. To begin with, in case of classification, our target variable needs to be binary or nominal and not ordinal. If our target variable was continuous then we will need to build a regression tree. The next step is for us to set out the goal of what we are trying to accomplish. In the case of a classification tree we will try to minimize classification error (or maximize accuracy) of the model. In case of a regression tree we may want to minimize least squares deviation (a measure closely related to variance).
Once we have set a goal for ourselves we need to think of the possible ways to accomplish this goal. First thing to notice is that it is not feasible to search through all the permutations and combinations of the possible partitions of the data based on all features and their possible split values. Therefore, we need a systematic way to decide on ways to split the data. Most tree algorithms use a greedy approach. This means that we first find an ‘optimal’ feature to split the data on and we keep that split for the rest of the tree construction process. This is a greedy method in the sense that we never go back to see if it is actually an optimal split once we further partitioned the data. This is done so that computation will be manageable.
With above in mind we can begin by first assigning all the data to the root node. The class label assigned to that node is the majority class. This means that if our dataset has 100 observations and 40 belong to class A while 60 belong to class B, then we will assign class B to the root node. This simply means that if we have no additional information about the data except the class counts then we will guess the majority class. In the Titanic data set, if we didn’t have any information about any particular passenger and wanted to guess if he/she survived then we would guess NO since in this data set 62% of passengers died.
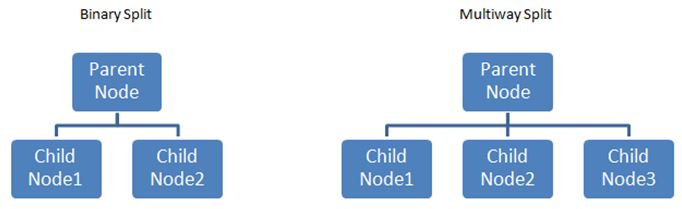
We can obviously improve on the above crude estimate by trying to split the data on one of the features that we deemed to be important for inclusion. The CART algorithm makes binary splits, that is, from each parent node there can be only two branches. Other algorithms allow for multiple splits.
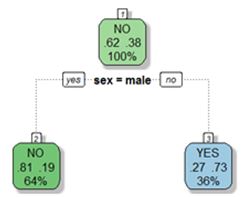
To tackle the problem systematically we need a measure of what is a good split. In either classification or regression problems what we really need to do is to minimize some measure of impurity in the subsets of data after each split. That is, once we split our sample into two (in case of binary splits), we want the data in each particular child nodes to be similar (ie pure). In our Titanic example, we started out with 62% of the individuals perishing in the sinking. After we split the data into two subsets based on gender, we ended up with less impurity in each node. In case of males we have 81% who did not survive. Meanwhile, in the subset of females, we have 73% that survived.
Splitting Criteria:
There are many ways to measure impurity. The two most commonly used impurity measures in classification trees are:
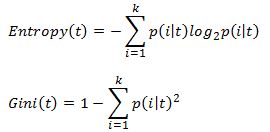
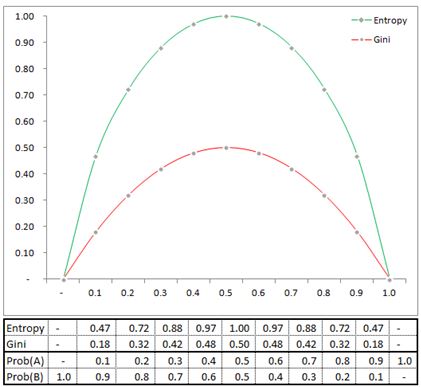
where k is the number of classes, and p(i|t) is the probability of class i at node t. The probability is simply the fraction of observations that belong to class i at node t. The Gini coefficient is used in the CART algorithm while Entropy is used in ID3, C4.5 and C5 algorithms.
In our parent node we have 62% of the observations belonging to class NO and 38% of the observations belonging to class YES. Therefore the Gini index is .4712 [1- .62^2-.38^2]. Lets pause for a second and see why Entropy or Gini are used. Notice that both are symmetric functions, if frequency of class A dominates or class B dominates both Entropy and Gini are low. When there is maximum impurity (ie 50% of data belongs to class A and the other 50% belongs to class B) then both Entropy and Gini are at their maximum.
After we have our impurity measure, we can test which split on a particular feature delivers the biggest reduction of total impurity. To do so, we will calculate the impurity measure of a node before any split, and compare it to an impurity measure after the split for each feature. Data partition that delivers the largest reduction in impurity will be selected. To calculate the reduction in impurity we can define Gain as:
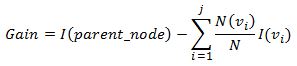

Above equation calculates a weighted average impurity of the split. The weights are based on the proportion of observations that are allocated to a child node.
Since impurity for a parent node is the same, we simply find a split that has the smallest impurity measure after a split (second term in the Gain equation).
Calculation Examples:
So now that we know that we are trying to reduce impurity in a data set with each split, and we know how to calculate this gain, let’s see how the computation is done on a stylized example. Below is a table with some fictitious data. We are trying to classify an individual as believing in global warming or not based on two features. The first feature measures the IQ of an individual (continuous variable) and the second feature is a binary feature that is coded as YES if an individual owns a tinfoil hat and NO otherwise (https://en.wikipedia.org/wiki/Tin_foil_hat). I also present a chart to the right of the data set.
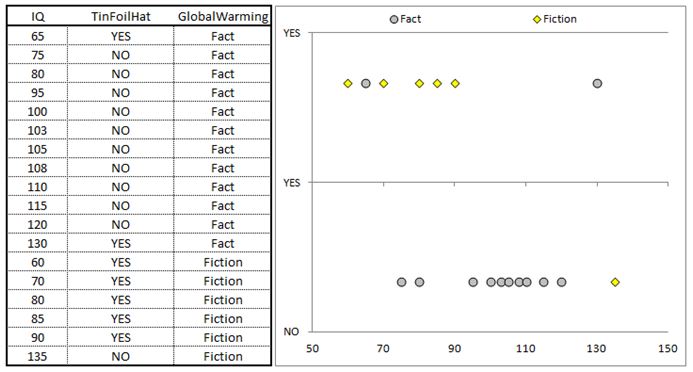
When building a tree we are basically attempting to find decision boundaries that will isolate clusters of data. This point will become apparent as we work through the example.
So the first thing we need to do is calculate the Gini index of the root node. In total we have 18 observations. We have 12 individuals that believe in global warming and 6 that do not. Therefore, p(Fact)=.67% [12/18] and p(Fiction) = .33 [6/18]. Remembering our Gini index formula we have Gini(root_node)=.444 [1-.67^2-.33^2].
Now we need to evaluate impurity of the data after a split. Lets first consider the binary feature. If we were to split the data based on whether or not an individual owns a tinfoil hat we end up with two subsets.
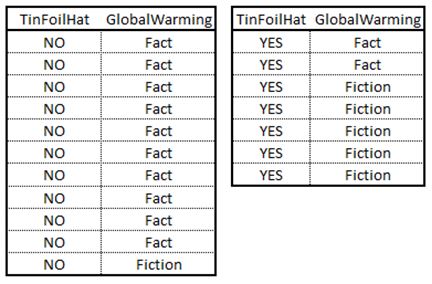
For the subset of individuals that do not own a tinfoil hat we have a total of 11 observations. Ten believe in global warming while one doesn’t. The probabilities are 91% in the first case [10/11] and 9% [1/11] in the latter. Therefore the Gini index for this subset is .165 [1-.91^2-.09^2]. Moving on to the second subset we have 7 observations. Here we have p(Fact) is 29% [2/7] while p(Fiction) is 71% [5/7]. This results in a Gini index of .490 [1-.29^2-.71^2]. The weighted average Gini index for this split is .291 [11/18*.165+7/18*.49]. Therefore, if we are to use this feature as a split we can achieve a Gain of .153 [.444-.291].
The calculations for a continuous variable follows the same idea but are a little more involved. First we sort the data in an ascending order.
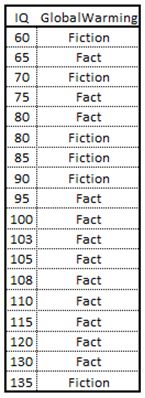
We then consider splits in between each IQ value. Below are the calculations for our data:
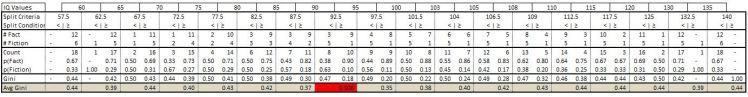
I will go through the calculations for one of the candidate splits. For example we consider a split of the data based on IQ being less than 92.5. In that case there are 5 individuals who believe global warming is fiction versus 3 that believe it is a fact. This produces a Gini index of .47. For the other subset of data that contains all individuals who’s IQ are above 92.5 we have 9 that believe global warming is a fact while only 1 believes its fiction. On this subset we get a Gini index of .18. Therefore the average Gini
index is equal to .31. Therefore our Gain calculation results in .136 [.444-.308]. This means that the Gain (ie impurity reduction) is larger by splitting the data based on the TinFoilHat feature.
Now we have one root node and two child nodes.
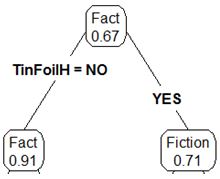
We can then repeat the procedure that I mentioned above by attempting to further subset the data for each individual child node. After repeating one more round of the calculations we introduce more depth to the tree. Below is an example of the final tree.
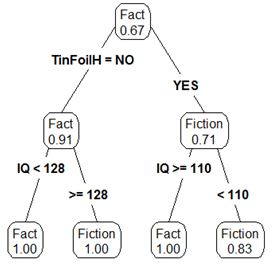
Lets now come back to the graph of the data. As I mentioned, we were effectively looking for a decision boundary. After performing the calculations we can see this more clearly. Our first split of the data introduced a split as seen below:
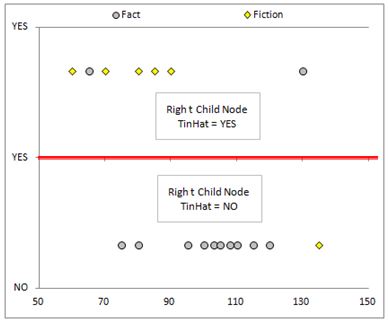
After splitting the data again, we introduced more decision boundaries.
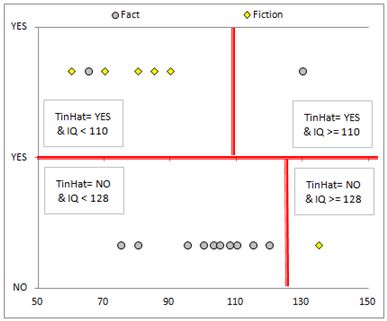
The Algorithm:
Let’s review quickly what we have established so far. We decided to use a greedy algorithm that performs a binary split of the data into two subsets. The feature, and the value at which to split the feature is decided based on purity gain criteria. The gain in purity is measured using the Gini index. After performing the initial split, we continue the process for each child node. The process continues until a stopping criteria is met. Obvious stopping criteria includes cases when all the observations in a node are the same (ie a pure node). Another criteria is when all the feature values are the same (ie we cannot split the feature anymore. Yet another criteria is when some minimum value of observations are allocated to a particular node. An example would be to stop splitting the data at a node when it has 10 observations or less. Finally, we can stop the algorithm when some maximum depth of a tree has been reached. For instance, if we don’t want a deep tree, we can stop the algorithm when we have a tree of depth 10.
After we generate a classification tree we can now use it to classify new observations. If we take a sample case and follow it to a leaf (terminal) node we assign the majority class in that node to the sample case. What I mean is that if we arrive at a terminal node and there are X observations belonging to class A and Y observations that belong to class B and if X>Y then we will assign class A to our sample case. If X<Y then we will obviously assign class B to the sample case.
Tree Pruning:
An obvious problem that we will almost surely encounter if we use above algorithm is that we will overfit the data. Generating bushy trees (ie trees with large depth) will leave us with a model that has high variance. If we were to get more sample data and then refit the model then we are likely to get a very different tree. To deal with this issue we can use a tree pruning method to reduce the variance of the model in exchange for increased bias. I can see that you are probably wondering why not avoid this issue by setting a small max depth stopping criteria in the main algorithm. The reason is that we can build a model that fits the data and generalizes much better if we use pruning.
If you have read my previous posts on Ridge and Lasso model then you should already have some idea of how we can introduce bias into the model to reduce the variance. In CART this is achieved via complexity parameter that is selected using a cross-fold validation technique. More specifically we calculate a Cost Complexity value for a tree:

The idea of CP is similar to the shrinkage (penalty) parameter lambda that we used in Ridge and Lasso regression models. In case of a classification tree we simply add a penalty for having more terminal nodes. So a model with alpha set to 0 will be a bushy tree. As we increase alpha we decrease the tree’s depth. As with all hyper-parameters , we select a value for alpha that minimizes misclassification on a cross-validation run.
Back to Titanic Example (in R):
I will jump straight into the model without any exploratory data analysis since the objective is to show you the basic steps of fitting a classification tree. Below is a printout of some of the data.
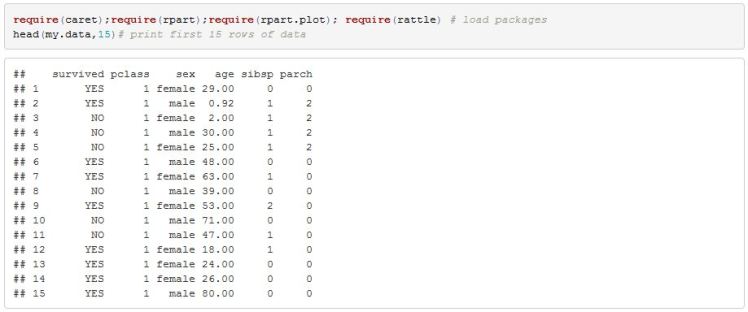
In order for us to assess the quality of the final model I will split the data into a training and testing set. The training set will contain 80% of the data. Splitting is done so that class balance is maintained (ie same proportion of survivors and non survivors is maintained across both sets of data).
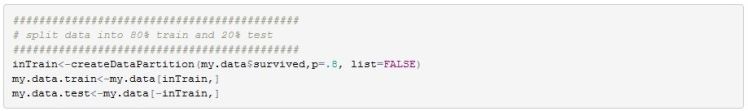
We can use R’s rpart package to fit a bushy model and also retrieve the optimal CP value
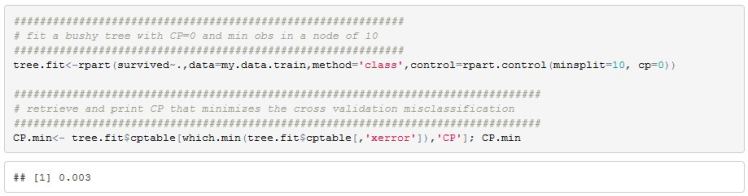
After creating an rpart object we can check the cross validation classification error across different CP values
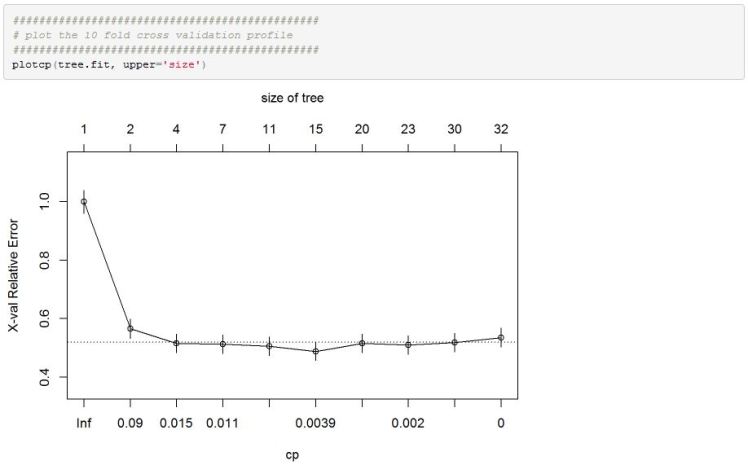
We can see that model performance measure flattens out past some initial restrictively large CP values.
Once we have settled on an optimal CP value of .003 we can plot this tree
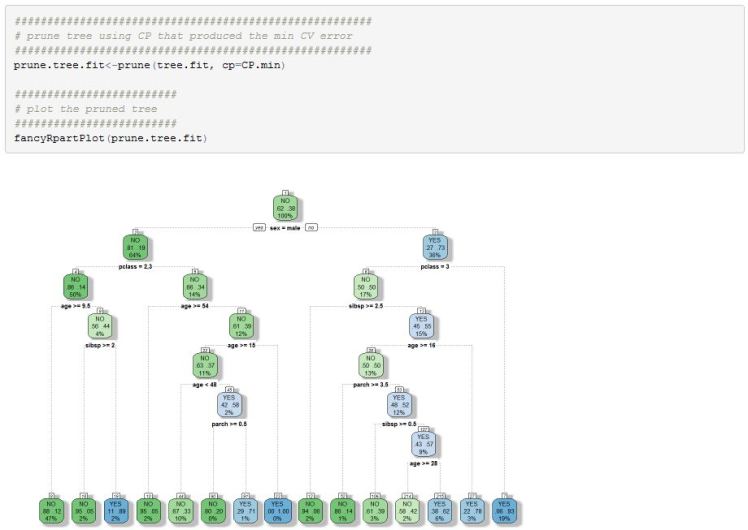
We can now use this model on our test data to get an estimate on how well it generalizes.
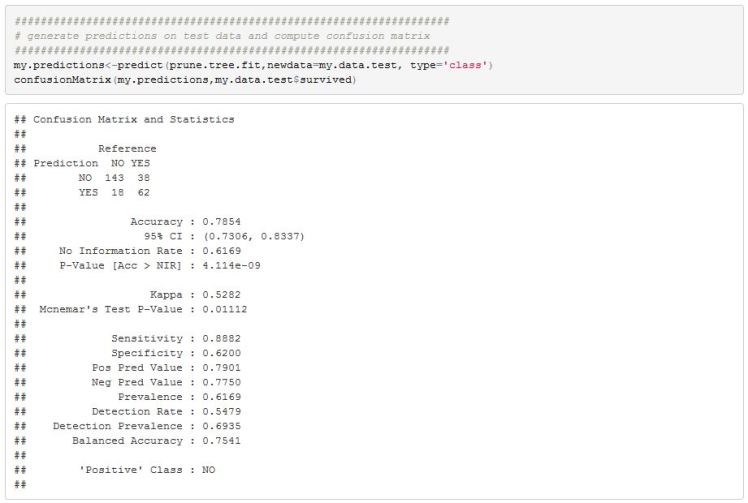
The final model has an accuracy rate of 79%. Not great but not bad either. There is plenty we can try to do to improve on this simple model but I think I will leave things here. Hopefully I was able to effectively convey the steps involved in generating a classification tree.
Couple Final Word:
This blog has been a little too long so I will briefly mention some important points.
- One additional benefit of tree models is that they can handle missing value using something called surrogate splits.
- Trees have been around for a long time and are not particularly great at prediction. There are bagging methods that improve predictive performance of trees at the expense of lost interpretability. Random forest models are an example and are very popular. Boosting methods also exist and are fun to play with. Gradient boosting machines are an example of boosting methods. There are excellent packages in R to build random forest and gbm models.
- I did not give credit to authors that I sourced for the blog but they are all mentioned at the bottom of the blog. All are excellent and strongly recommended.
- I have not read much of the literature on trees’ use in finance but there are published papers on the use of trees for stock portfolio construction. Just mentioning it for those who are interested.
Some Useful Resources:
1) Introduction to Data Mining has an excellent chapter on trees. I highly recommend this source. I found the presentation very clear. http://www-users.cs.umn.edu/~kumar/dmbook/index.php
2) rpart writeup that is excellent http://www.mayo.edu/research/documents/biostat-61pdf/doc-10026699?_ga=1.78265300.1735958202.1452508562
3) Creators of CART algorithm wrote an excellent book that I used as reference but have not gone through the entire book. Leo Breiman, Jerome Friedman, Richart Olshen, Charles Stone “Classification and Regression Trees” http://www.amazon.com/Classification-Regression-Wadsworth-Statistics-Probability/dp/0412048418
4) Even though I didn’t use the charts here an excellent discussion of prp function in R to plot decision trees http://www.milbo.org/rpart-plot/prp.pdf
5) Quick and excellent example of rpart functions in R http://www.statmethods.net/advstats/cart.html
6) Max Kuhn and Kjell Johnson discuss many important decisions that need to be made to build effective predictive models (including regression and classification trees) http://appliedpredictivemodeling.com/
7) Introduction to Statistical Learning by James, Witten, Hastie, and Tibshirani is an excellent book with a light discussion of trees and their estimation in R http://www-bcf.usc.edu/~gareth/ISL/